Casein, the major milk protein, is a macromolecule composed of amino acid subunits linked together by peptide bonds (CO—NH). It makes around 85% of the protein found in milk as well as the white color of milk.
Casein is way too large to enter the cell membrane. Before their assimilation into the cell, proteins must undergo step-by-step degradation into peptones, polypeptides, dipeptides, and ultimately into their building blocks, amino acids.
These are mediated by extracellular enzymes called proteases. The function of these proteases is to cleave the peptide bond CO–NH by introducing water into the molecule. The reaction then liberates the amino acids which are low-molecular in weight which can be transported through the cell membrane for use in the synthesis of structural and functional cellular proteins.
Objectives
- To determine the ability of the organism to degrade the casein protein.
- To differentiate the organism on the basis of production of exoenzyme proteinase (caseinase)
Principle
Some microorganisms have the ability to degrade the casein protein by producing proteolytic exoenzyme, called proteinase (caseinase). For demonstration of such an activity, in the lab, milk agar is used. The medium is composed of nutrient agar supplemented with milk that contains the protein substrate casein. Similar to other proteins, milk protein is a colloidal suspension that gives the medium its color and opacity because it deflects light rays rather than transmitting them.
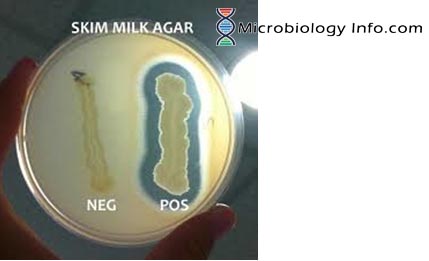
Following inoculation and incubation of the agar plate cultures, organisms secreting proteases will exhibit a zone of proteolysis, which is demonstrated by a clear area surrounding the bacterial growth. This loss of opacity is the result of a hydrolytic reaction yielding soluble, non-colloidal amino acids, and it represents a positive reaction.
In the absence of protease activity, the medium surrounding the growth of the organism remains opaque, which is a negative reaction.
Media
Skim milk agar
SM powder 28.0gm/L, Tryptone 5.0gm/L, Yeast extract 2.50gm/L, Dextrose (Glucose) 1.0gm/L, Agar 15.0gm/L, Final pH ( at 25°C)
Method
- Inoculate the organism on the plate either a straight line or a zig-zag.
- Incubate the plate at 25°C or 37°C.
- Examine the milk agar plate cultures for the presence or absence of a clear area, or zone of proteolysis, surrounding the growth of each of the bacterial test organisms.
Result Interpretation
- Positive Test: Clearing is observed around and/or beneath colony growth (hydrolysis).
- Negative Test: No clearing is observed around and/or beneath the inoculum.
Uses
- Helpful in identifying bacteria that grow in milk
- Differentiating among Enterobacteriaceae, Bacillaceae, and several other families.
- For the differentiation of aerobic actinomycetes based on casein proteolysis.
- Identification of organisms that can hydrolyse casein, such as Streptomyces, Pseudomonas , and Actinomadura
Limitations
- It is recommended that biochemical, immunological, molecular, or mass spectrometry testing be performed on colonies from pure culture for complete identification.
References
- Cappuccino J.G. and Sherman N. 2008. Microbiology: A Laboratory Manual, 8th ed. Pearson Benjamin Cummings, San Francisco, CA, USA.
- Aneja K.R. 2003. Experiments in Microbiology, Plant Pathology and Biotechnology, fourth revised edition, New Age International (P) limited, Ansari road, Daryaganj, New Delhi-110002.
- https://catalog.hardydiagnostics.com/cp_prod/Content/hugo/CaseinAgar.html
- Brown A.E. 2009. Benson’s Microbiological Applications: Laboratory Manual in General Microbiology, 11th ed. McGraw-Hill Companies, New York, NY, USA.
- https://homepages.wmich.edu/~rossbach/bios312/LabProcedures/Casein%20hydrolysis%20test%20results.html
- delrio.dcccd.edu/jreynolds/microbiology/2421/lab_manual/casein.pdf
- https://jcp.bmj.com/content/jclinpath/25/12/1083.full.pdf

This website has been very helpful in obtaining information on Biochemical Activities of Microbes.
A five star rating for that.