Cetrimide Agar is a selective and differential medium used for the isolation and identification of Pseudomonas aeruginosa from clinical and non-clinical specimens. Cetrimide is the selective agent and inhibits most bacteria by acting as a detergent ( Cetyltrimethylammonium bromide, a quaternary ammonium, cationic detergent).
It is also known as Pseudomonas Cetrimide Agar or Pseudosel Agar.
Composition of Cetrimide Agar
| Ingredients | In gm/Litre |
|---|---|
| Pancreatic Digest of Gelatin | 20.0 gm |
| Potassium Sulfate | 10.0 gm |
| Magnesium Chloride | 1.4 gm |
| Cetyltrimethylammonium Bromide | 0.3 gm |
| Glycerine | 10.0 ml |
| Agar | 13.6 gm |
| Distilled Water | 1000 ml |
Final pH 7.2 +/- 0.2 at 25 degrees C
Principle of Cetrimide Agar
Cetrimide Agar is used as a selective medium for the isolation of Pseudomonas aeruginosa from pus, sputum and
drains, etc. Pseudomonas aeruginosa produces a number of water soluble iron chelators, including the yellow-green or yellow-brown fluorescent pyoverdin. When pyoverdin combines with the blue water-soluble pyocyanin, the bright green color characteristic of Pseudomonas aeruginosa is created.
Pancreatic digest of gelatin provide necessary nutrients for P. aeruginosa such as nitrogen, vitamins, and carbon.
The addition of magnesium chloride and potassium sulphate stimulates pyocyanin and pyoverdin (fluorescein) production.
Cetyltrimethylammonium bromide (Cetrimide) is the selective agent and inhibits most bacteria by acting as a detergent. When in contact with bacteria, causes the release of nitrogen and phosphorous from the bacterial cell other than Pseudomonas aeruginosa.
Glycerol is supplemented as a source of carbon.
Agar is the solidifying agent.
Uses of Cetrimide Agar
- It is primarily used for the selective isolation and presumptive identification of Pseudomonas aeruginosa
from clinical and nonclinical specimens. - It is also used for determining the ability of an organism to produce fluorescein and pyocyanin (Antibiotica)
Preparation of Cetrimide Agar
- Add 45.3 gm of the medium in 1 litre of distilled water.
- Add 10ml of glycerol and boil to dissolve completely.
- Sterilize by autoclaving at 121°C for 15 minutes.
- Cool the medium to approximately 50°C and pour into sterile Petri dishes.
Interpretation of Results on Cetrimide Agar
The presence of growth is indicative of a positive reaction. Examine colonies under short wavelength (254nm) ultraviolet light for the presence of fluorescein. Visual examination may also reveal the typical yellow-green to blue color which indicates the production of pyocyanin. Both pyocyanin and fluorescein are typically produced by strains of P. aeruginosa.
A negative reaction is denoted by no growth.
Colony Morphology on Cetrimide Agar
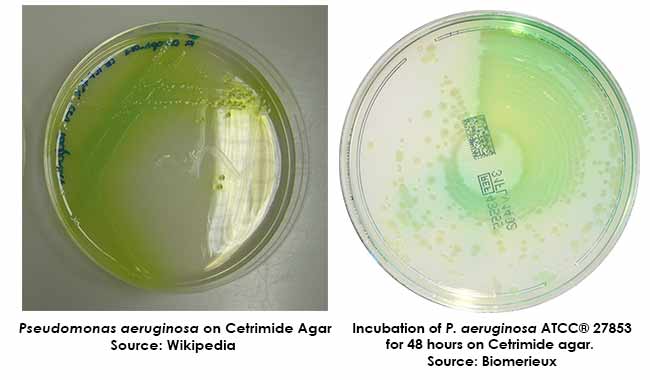
Quality Control on Cetrimide Agar
Pseudomonas aeruginosa ATCC 9027 – Yellow-green to blue colonies.
Escherichia coli ATCC 8739 – Partial to complete inhibition. No Pigmentations.
Limitation of Cetrimide Agar
- Occasionally some enterics will exhibit a slight yellowing of the medium; however, this coloration is easily
distinguished from fluorescein production because this yellowing does not fluoresce. - Some non-fermenters and some aerobic spores formers may exhibit a water-soluble tan to brown pigmentation on this medium. Serratia strains may exhibit a pink pigmentation.
- Studies of Lowbury and Collins showed P. aeruginosa can lose its fluorescence under UV if the cultures are left at room temperature for a short time. Fluorescence reappears when plates are re-incubated.
- Further tests are necessary for confirmation of P. aeruginosa.
Similar Posts:
- Cetrimide Test – Principle, Procedure, Uses and Interpretation
- List of culture media used in microbiology with their uses
- Thiosulfate-Citrate-Bile Salts-Sucrose (TCBS) Agar- Composition, Principle, Uses, Preparation and Colony Morphology
- Xylose Lysine Deoxycholate (XLD) Agar- Principle, Uses, Composition, Preparation and Colony Characteristics

We currently utilise Cetrimide agar as part of our routine water monitoring (alongside testing on R2A). We perform membrane filtration and transfer the filter onto Cetrimide agar before incubating this at 35-37°C. A few months ago we had an out of trend result where growth was observed on the filter of the cetrimide plate. When the ID came back it was identified as normal skin flora (mainly staphylococcus). Given the properties of CET agar, this was investigated further as growth of staphylococcus was thought to be inhibited by this agar (although EP/USP does not call for S.aureus testing as part of QC).
The investigative study we performed involved directly spiking the CET agar with S.aureus and also performing the membrane filtration method spiking the final rinse with S.aureus. We found that when the plate is spiked directly, no growth of S.aureus is observed, however when the rinse for the membrane filtration method is spiked S.aureus growth was observed on the filter.
It appears to be that when utilising a membrane filter on CET agar that the bacteria on the filter are protected from the inhibitory properties of the agar. I was wondering if anyone has any further advice to offer on this scenario? Or if any additional relevant studies have been performed? From a quick online search I can see that other outlets utilise Cetrimide agar and membrane filtration but I can’t find much data around the issue we have encountered. The testing is for information only so we have no major concerns, but we don’t want to be performing a test if it is of no value at all.
Can ps.aeruginosa viable for 12days (288hrs)of extended incubation on cetrimide agar
I have question regarding Cetrimide agar. We are doing water testing for the presence of P.aeruginosa. We noticed that water is contaminated by Pseudomonas aeruginosa at the outlet of a water system, after draining the system, we wanted to identify the source of the contamination by doing surface swabbing, but the tests didn’t reveal the presence of pseudomonas. Is it possible to increase the sensibility (or possibility to recover the pseudo) by increasing the incubation time, eg 4-5 days of incubation ?
You may need to do an enrichment step before the plate, after collecting the swab do an enrichment on TSB for 18-14 hrs @ 30-35C, then streak onto CET or other media you need to. The enrichment will help with the recovery of stress microorganisms
how to produce yellow zone by staphylococcus aureus? and
how to produce gas by some organisms?
What is the main role of glycerol in cetrimide agar
It serves as a carbon supplement
what happen if we don’t add glycerol in centrimide? Is that It will affect the result?
Will Pseudomonas fluorescens grow on cetrimide agar?
I have question regarding Cetrimide agar. We are doing soil testing for the presence of P.spp .
There are no colonies on the petri medium cetrimid
While soil that has a high population of Pseudomonas
Soil contains a variety of organisms. It is possible that in the soil sample the high population of Pseudomonas is of different species. The Cetrimide plate is specifically for Pseudomonas aeruginosa organisms only as it is a selective in nature. Differential in nature since it allows the growth of P.aeruginosa specifically and not any other Pseudomonas species
Maybe there was error in your serial dilutions
I have question regarding Cetrimide agar. We are doing water testing for the presence of P.aeroginosa.
There are no colonies on the membrane filter, however there is formation of green fluorescein under the filter that glows under UV. Is this indicating the positive results of P.aeroginosa? If so, why there are no colonies on the filter?
Membrane filter paper have no count means it is a qualitative method
Where as pesudomonas bacteria not detected on the filter paper it is a qualitative method it require the enrichment and differential media like cetramide agar