Polymerase Chain Reaction (PCR) is a powerful method for amplifying particular segments of DNA, distinct from cloning and propagation within the host cell. This procedure is carried out entirely biochemically, that is, in vitro. PCR was invented by Kary Mullis in 1983. He shared the Nobel Prize in chemistry with Michael Smith in 1993.
Principle of PCR
PCR uses the enzyme DNA polymerase that directs the synthesis of DNA from deoxynucleotide substrates on a single-stranded DNA template. DNA polymerase adds nucleotides to the 3` end of a custom-designed oligonucleotide when it is annealed to a longer template DNA. Thus, if a synthetic oligonucleotide is annealed to a single-stranded template that contains a region complementary to the oligonucleotide, DNA polymerase can use the oligonucleotide as a primer and elongate its 3` end to generate an extended region of double stranded DNA.
Procedure/Steps of PCR
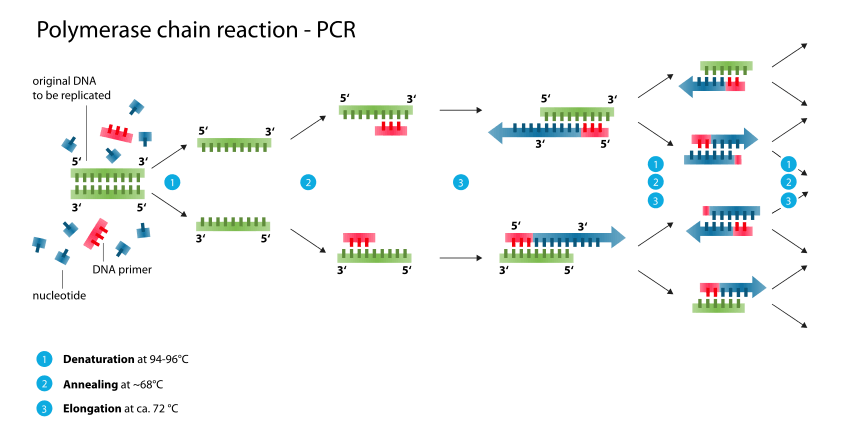
By Enzoklop (Own work) via Wikimedia Commons
1. Denaturation
The DNA template is heated to 94° C. This breaks the weak hydrogen bonds that hold DNA strands together in a helix, allowing the strands to separate creating single stranded DNA.
2. Annealing
The mixture is cooled to anywhere from 50-70° C. This allows the primers to bind (anneal) to their complementary sequence in the template DNA.
3. Extension
The reaction is then heated to 72° C, the optimal temperature for DNA polymerase to act. DNA polymerase extends the primers, adding nucleotides onto the primer in a sequential manner, using the target DNA as a template.
With one cycle, a single segment of double-stranded DNA template is amplified into two separate pieces of double-stranded DNA. These two pieces are then available for amplification in the next cycle. As the cycles are repeated, more and more copies are generated and the number of copies of the template is increased exponentially.
Types of PCR
- Real-time PCR
- Quantitative real time PCR (Q-RT PCR)
- Reverse Transcriptase PCR (RT-PCR)
- Multiplex PCR
- Nested PCR
- Long-range PCR
- Single-cell PCR
- Fast-cycling PCR
- Methylation-specific PCR (MSP)
- Hot start PCR
- High-fidelity PCR
- In situ PCR
- Variable Number of Tandem Repeats (VNTR) PCR
- Asymmetric PCR
- Repetitive sequence-based PCR
- Overlap extension PCR
- Assemble PCR
- Intersequence-specific PCR(ISSR)
- Ligation-mediated PCR
- Methylation –specifin PCR
- Miniprimer PCR
- Solid phase PCR
- Touch down PCR, etc
Applications of PCR
- PCR is used in analyzing clinical specimens for the presence of infectious agents, including HIV, hepatitis, malaria, anthrax, etc.
- PCR can provide information on a patient’s prognosis, and predict response or resistance to therapy. Many cancers are characterized by small mutations in certain genes, and this is what PCR is employed to identify.
- PCR is used in the analysis of mutations that occur in many genetic diseases (e.g. cystic fibrosis, sickle cell anaemia, phenylketonuria, muscular dystrophy).
- PCR is also used in forensics laboratories and is especially useful because only a tiny amount of original DNA is required, for example, sufficient DNA can be obtained from a droplet of blood or a single hair.
- PCR is an essential technique in cloning procedure which allows generation of large amounts of pure DNA from tiny amount of template strand and further study of a particular gene.
- The Human Genome Project (HGP) for determining the sequence of the 3 billion base pairs in the human genome, relied heavily on PCR.
- PCR has been used to identify and to explore relationships among species in the field of evolutionary biology. In anthropology, it is also used to understand the ancient human migration patterns. In archaeology, it has been used to spot the ancient human race. PCR commonly used by Paleontologists to amplify DNA from extinct species or cryopreserved fossils of millions years and thus can be further studied to elucidate on.

What about the tetra ARMS PCR technique?
i just love the way that PCR is explained. its to easy to understand. thanks for this
Best principle of PCR ever explained. Thank you !!!!
I was started working on PCR after finishing my Master, loved the way it produce the accurate and fast result. I was running 3 to 4 test at a time and it gives amazing result. With the help of PCR result it become easy to find how much infection in the blood and based on infection the Specialist doctor can treat the patient and the strength of medicine can be given to cure fastest.
Thanks to those people who found this way of doing.
Hi Sir!One important question.
How the PCR is working in microbial diagonistics in medical.
specific genetic regions of different microbs are detected and amplified with the help of known primers.
Dear All, I am a medical Microbiologist. I’m interested in Post doctorate fellowship research into genetic studies of Multi-Drug Resistant Tuberculosis. Can somebody link me up. Thanks
how can this polymerase chain reaction be applied in the detection of zika virus in the lab ?
what are u doing explain ur work lil briefly
simply PCR work in 3 steps they are Denaturation where the dubble strand DNA break into single strand. Its temperature is 94 degree Celsius, than annealing temperature where primer bind it temp generally 50-70 degree Celsius and finally extension where the Taq polymerase act on the DNA and primer joining them with hydrogen bonding.
simply PCR is the machine which give control temperature, where we prepare our reaction mixture which contain 1.Taq buffer 2.Taq polymerase 3. dNTP which provide nucleotide for the new strands 4.Forward and reverse primer for The reverse primer is designed to attach to the complementary strand to synthesize DNA in the reverse direction — towards the forward primer. The primers are added to PCR experiments to initiate the process of replication by providing the initial nucleotides to the new strand. 5. Template DNA 6. HPLC graded water.
I am working on a bacteria specie with intention of isolating it from different sources and comparing their antibiotic resistance plasmid content and other characteristics.How do I determine the right primers to use for my work?
First i most comment your effort you’re doing great. I am writting to see if anyone can link me to a reputable university where i can run P.hD in molecular microbiology with MRSA as my microorganism of interest. I worked on it in my Masters and i want to continue with it. Am waiting for any assistance one can render.
wat s d role of Thermus aquaticus in PCR and how does it works?
It’s a thermo stable bacterium (Thermus aguatics ) to maintain a temperature duringPCR
An enzyme DNA polymerase is an indispensable tool for DNA replication, we all knows that enzymes are temperature sensitive, Taq DNA is DNA polymerase that’s isolated from bacteria Thermus aquaticus, This enzyme can withstand different temperatures 92,54, 72 degrees..
If during amplification of ur desired DNA (wether Replication or PCR) ur polymerase became inactivated by heat, then the process adding nucleotides and subsequent elongation stops. Therefore Taq DNA is the suitable enzyme to used.
thermus aquaticus is a bacterial specie from which the polymerase enzyme can be obtaines which is use for polymerization in pcr
The taq polymerase used for PCR is from Thermus Aquaticus. And it does not “maintain a tempreature during PCR” but, tolerates and survives in high tempreature, unlike most other bacteria. PCR uses high tempreatures and these polymerases hence work effectively.
I love pcr
Awesome info. Thanks!
It would be nice if you share the distinctive futures of the specific types of PCR too.
Kind regards
Hi Sir!One important question.
How the PCR is working in microbial diagonistics in medical.
I need a perfect to present a seminar so plzzz reply me “AS SOON AS POSSIBLE”.
“IAM EXPECTING MORE FROM THIS WEBSITE”
1st analyse the microbe DNA and get sequence and then identified the microbe specious then proceed for antimicrobial methods.
dr how can downlaod the books of immunology?
The multiplex endpoint PCR technology offers a number of potential advantages, results are available in a matter of hours rather than days, the extreme sensibility facilitates detection of even minutes the amounts of pathogen DNA in clinical samples.
While microbiological culture is likely to remain a gold standard for infection diagnosis, there is growing interest at the potential of PCR technology to provide early, time critical information based on detection and recognition of bacterial or fungal pathogen DNA. By this new modern method, the results are chosen in perquisite for giving adequate antibiotics treatment as early as possible in order to improve the standard of care.
Dr. Aurel